Two easy additions to strain engineering DBTL workflows to better understand and protect your microbial assets
By Dr Reuben Carr, Technology Commercialisation Manager, NCIMB Ltd.
Acquiring precision genetic alignment between strain design and biological reality can be a challenge for microbial strain engineers. NCIMB’s technology commercialisation manager Dr Reuben Carr considers what is at stake, and the steps that can be taken to tackle this important issue.
Precision engineered microbes provide sustainable and low carbon routes to the production of essential chemical feedstocks. They contribute billions to the global economy and help achieve industrial net zero targets. Whilst some precision engineered microbes are already put to work in large scale fermentation processes – for example to produce L-lysine and other proteinogenic amino acids, which find application in use as nutritional supplements, flavour enhancers and in pharmaceutical manufacturing; there is scope for many more important products to be manufactured using fermentation.
Design-Build-Test-Learn (DBTL) using precision engineering of microbes has become the standard approach to bioengineering to obtain new phenotypes for the desired or improved industrial application. This approach has brought significant benefits, including the ability to automate and scale. However, the inherent complexity of living biological organisms means that challenges inevitably remain. These include strain stability, genetic drift, and demonstration of the nucleotide level alignment of strain genetic design assumption and the biological reality of the engineered strain’s genetic makeup – in other words is the strain produced exactly what you think it is? This is not just a technical challenge; it also brings regulatory and quality risks in delivering products to market.
Why biological reality often deviates from strain design assumptions
Deviations from the strain design assumptions during genomic engineering can arise for several reasons:
While planned changes to the genome are typically well tracked and monitored, for example by PCR amplification of the modified region, there is a non-zero probability that unplanned changes in the genome also occur. These can include off-target integration of donor DNA into the genome, transposon migration of integrated elements. These unplanned and unscheduled modifications compromise the precision of the strain design output.
These aberrations of the genome from the parental whole genome sequence input can compromise further genomic engineering efforts on DBTL cycles – particularly where the efficiency of the tools to manipulate the genome rely on sequence complementarity and homology with precision molecular based tools such as Flp–FRT, homologous recombination, integrative plasmids, CRISPR and others. Any base pair mismatches between designed sequences and genome target sites quickly escalates the failure rate for building the designed strain.
Another issue that arises as an inherent consequence of the DBTL approach is strain drift. Whenever a microbial strain is grown during strain engineering workflows, for example agar plating or liquid growth, there is potential for genetic drift. Each cell doubling is a roll of the DNA mutation dice, and even single-nucleotide changes can lead to editing failure, off-target effects, or general lack of predictability and control in genome modification. This negatively impacts internal workflow through remedial counter measures and can erode confidence in a strain’s genetic tractability for further DBTL cycles. The more DBTL cycles undertaken, the greater the risk of strain divergence from the parental strain.
Simple solution
Strain engineers often rely on the use of public domain third party genome sequence information to guide their precision genomic engineering efforts, but in my opinion, it is far better to understand the biological reality of the material you are working with by undertaking whole genome sequencing and sequence alignment on input and output strains of the DBTL cycle.
This valuable, but often overlooked step, would ensure that the strain engineers are fully informed throughout the process, preventing genomic tool manipulation misalignment, reducing DBTL failures in microbial strain engineering (saving costs), strengthening client IP, and supporting quality, and regulatory readiness to take the strain to market.
Whole genome sequencing and sequence alignment on each turn of the DBTL cycle is a valuable, but often overlooked step.
Monitoring the genome throughout the process increases confidence that cycles have resulted in successful outcomes and serves as a platform to provide greater predictability in future desired modifications.
Another straightforward, but not routinely undertaken step is preservation and storage of the genetically modified strains produced in each cycle. This would allow for a quick recovery should disaster strike during the project – allowing you to return to any point in the strain engineering programme, rather than starting from scratch.
Implementing robust and rigorous genetic alignments of your input/output engineered strains in combination with strain storage as best practice steps into workflows would support engineers and synthetic biologists in fully understanding and protecting core company assets.
How NCIMB can help
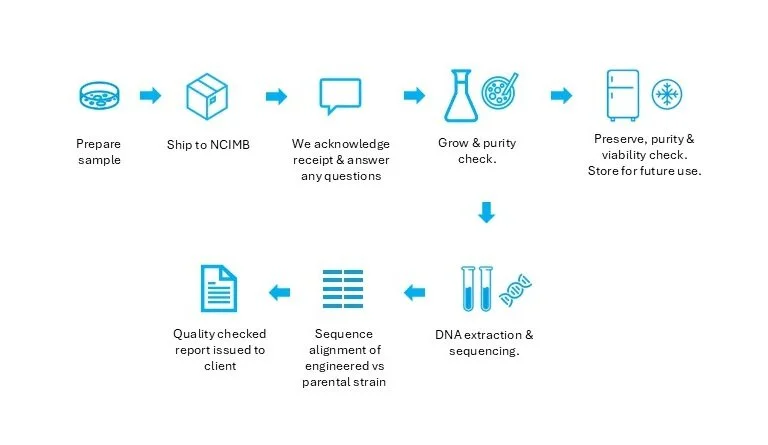
NCIMB provides an integrated, frictionless service to support companies working in microbial DBTL. The client simply sends the strain, and NCIMB manages everything from there on: this includes whole-genome sequencing, sequence alignment, preservation and storage, with chain-of-custody documentation.
Why not get in touch at enquiries@ncimb.com to learn more about how NCIMB can support your microbial engineering Design, Build, Test and Learn workflows? We are ready to receive your materials today.